Perform a series of checks to ensure that the local maps in each process still preserve map properties Let us run our usual tests to verify that the map data has been distributed correctly 1) Consistency check – row sum = 1.0 2) Disable conservation checks that require global column sums (and hence more communication/reduction) 3) Perform monotonicity check and ensure failures globally are same as serial case
72 NcError
error( NcError::silent_nonfatal );
76 NcVar *varRow = NULL, *varCol = NULL, *varS = NULL;
80 const int rootProc = 0;
85 const int nBufferSize = 64 * 1024;
86 const int nNNZBytes = 2 *
sizeof( int ) +
sizeof(
double );
87 const int nMaxEntries = nBufferSize / nNNZBytes;
88 int nA = 0, nB = 0, nVA = 0, nVB = 0, nS = 0;
89 double dTolerance = 1e-08;
91 int nprocs = 1,
rank = 0;
94 MPI_Comm_size( commW, &nprocs );
95 MPI_Comm_rank( commW, &
rank );
99 if(
rank == rootProc )
102 ncMap =
new NcFile(
inMapFilename.c_str(), NcFile::ReadOnly );
103 if( !ncMap->is_valid() )
105 _EXCEPTION1(
"Unable to open input map file \"%s\"",
inMapFilename.c_str() );
109 NcDim* dimNA = ncMap->get_dim(
"n_a" );
112 _EXCEPTIONT(
"Input map missing dimension \"n_a\"" );
117 NcDim* dimNB = ncMap->get_dim(
"n_b" );
120 _EXCEPTIONT(
"Input map missing dimension \"n_b\"" );
125 NcDim* dimNVA = ncMap->get_dim(
"nv_a" );
128 _EXCEPTIONT(
"Input map missing dimension \"nv_a\"" );
131 nVA = dimNVA->size();
133 NcDim* dimNVB = ncMap->get_dim(
"nv_b" );
136 _EXCEPTIONT(
"Input map missing dimension \"nv_b\"" );
139 nVB = dimNVB->size();
142 NcDim* dimNS = ncMap->get_dim(
"n_s" );
145 _EXCEPTION1(
"Map file \"%s\" does not contain dimension \"n_s\"",
inMapFilename.c_str() );
150 varRow = ncMap->get_var(
"row" );
153 _EXCEPTION1(
"Map file \"%s\" does not contain variable \"row\"",
inMapFilename.c_str() );
156 varCol = ncMap->get_var(
"col" );
159 _EXCEPTION1(
"Map file \"%s\" does not contain variable \"col\"",
inMapFilename.c_str() );
162 varS = ncMap->get_var(
"S" );
165 _EXCEPTION1(
"Map file \"%s\" does not contain variable \"S\"",
inMapFilename.c_str() );
170 int nEntriesRemaining = nS;
171 int nBufferedReads =
static_cast< int >( ceil( ( 1.0 * nS ) / ( 1.0 * nMaxEntries ) ) );
172 int runData[6] = { nA, nB, nVA, nVB, nS, nBufferedReads };
174 mpierr = MPI_Bcast( runData, 6, MPI_INT, rootProc, commW );
178 if(
rank != rootProc )
185 nBufferedReads = runData[5];
188 printf(
"Global parameters: nA = %d, nB = %d, nS = %d\n", nA, nB, nS );
190 printf(
"Global parameters: nA = %d, nB = %d, nS = %d\n", nA, nB, nS );
193 std::vector< int > rowOwnership( nprocs );
194 int nGRowPerPart = nB / nprocs;
195 int nGRowRemainder = nB % nprocs;
196 rowOwnership[0] = nGRowPerPart + nGRowRemainder;
197 for(
int ip = 1, roffset = rowOwnership[0]; ip < nprocs; ++ip )
199 roffset += nGRowPerPart;
200 rowOwnership[ip] = roffset;
205 SparseMatrix< double >& sparseMatrix = mapRemap.GetSparseMatrix();
208 if(
rank == rootProc )
210 printf(
"Parameters: nA=%d, nB=%d, nVA=%d, nVB=%d, nS=%d, nNNZBytes = %d, nBufferedReads = %d\n", nA, nB, nVA,
211 nVB, nS, nNNZBytes, nBufferedReads );
213 std::map< int, int > rowMap, colMap;
214 int rindexMax = 0, cindexMax = 0;
219 for(
int iRead = 0; iRead < nBufferedReads; ++iRead )
222 DataArray1D< int > vecRow;
223 DataArray1D< int > vecCol;
224 DataArray1D< double > vecS;
225 std::vector< std::vector< int > > dataPerProcess( nprocs );
226 std::vector< int > nDataPerProcess( nprocs, 0 );
227 for(
int ip = 0; ip < nprocs; ++ip )
228 dataPerProcess[ip].reserve( std::max( 100, nMaxEntries / nprocs ) );
231 if(
rank == rootProc )
234 int nLocSize = std::min( nEntriesRemaining,
static_cast< int >( ceil( nBufferSize * 1.0 / nNNZBytes ) ) );
236 printf(
"Reading file: elements %ld to %ld\n", offset, offset + nLocSize );
239 vecRow.Allocate( nLocSize );
240 vecCol.Allocate( nLocSize );
241 vecS.Allocate( nLocSize );
243 varRow->set_cur( (
long)( offset ) );
244 varRow->get( &( vecRow[0] ), nLocSize );
246 varCol->set_cur( (
long)( offset ) );
247 varCol->get( &( vecCol[0] ), nLocSize );
249 varS->set_cur( (
long)( offset ) );
250 varS->get( &( vecS[0] ), nLocSize );
253 for(
size_t i = 0; i < vecRow.GetRows(); i++ )
260 if( rowOwnership[0] > vecRow[i] )
264 for(
int ip = 1; ip < nprocs; ++ip )
266 if( rowOwnership[ip - 1] <= vecRow[i] && rowOwnership[ip] > vecRow[i] )
274 assert( pOwner >= 0 && pOwner < nprocs );
275 dataPerProcess[pOwner].push_back( i );
279 nEntriesRemaining -= nLocSize;
281 for(
int ip = 0; ip < nprocs; ++ip )
282 nDataPerProcess[ip] = dataPerProcess[ip].
size();
286 int nEntriesComm = 0;
287 mpierr = MPI_Scatter( nDataPerProcess.data(), 1, MPI_INT, &nEntriesComm, 1, MPI_INT, rootProc, commW );
291 if(
rank == rootProc )
294 std::vector< MPI_Request > cRequests;
295 std::vector< MPI_Status > cStats( 2 * nprocs - 2 );
296 cRequests.reserve( 2 * nprocs - 2 );
299 std::vector< std::vector< int > > dataRowColsAll( nprocs );
300 std::vector< std::vector< double > > dataEntriesAll( nprocs );
301 for(
int ip = 1; ip < nprocs; ++ip )
303 const int nDPP = nDataPerProcess[ip];
306 std::vector< int >& dataRowCols = dataRowColsAll[ip];
307 dataRowCols.resize( 2 * nDPP );
308 std::vector< double >& dataEntries = dataEntriesAll[ip];
309 dataEntries.resize( nDPP );
311 for(
int ij = 0; ij < nDPP; ++ij )
313 dataRowCols[ij * 2] = vecRow[dataPerProcess[ip][ij]];
314 dataRowCols[ij * 2 + 1] = vecCol[dataPerProcess[ip][ij]];
315 dataEntries[ij] = vecS[dataPerProcess[ip][ij]];
318 MPI_Request rcsend, dsend;
319 mpierr = MPI_Isend( dataRowCols.data(), 2 * nDPP, MPI_INT, ip, iRead * 1000, commW, &rcsend );
321 mpierr = MPI_Isend( dataEntries.data(), nDPP, MPI_DOUBLE, ip, iRead * 1000 + 1, commW, &dsend );
324 cRequests.push_back( rcsend );
325 cRequests.push_back( dsend );
331 int rindex = 0, cindex = 0;
332 assert( dataPerProcess[0].
size() - nEntriesComm == 0 );
333 for(
int i = 0; i < nEntriesComm; ++i )
335 const int& vecRowValue = vecRow[dataPerProcess[0][i]];
336 const int& vecColValue = vecCol[dataPerProcess[0][i]];
338 std::map< int, int >::iterator riter = rowMap.find( vecRowValue );
339 if( riter == rowMap.end() )
341 rowMap[vecRowValue] = rindexMax;
346 rindex = riter->second;
348 std::map< int, int >::iterator citer = colMap.find( vecColValue );
349 if( citer == colMap.end() )
351 colMap[vecColValue] = cindexMax;
356 cindex = citer->second;
358 sparseMatrix( rindex, cindex ) = vecS[i];
362 mpierr = MPI_Waitall( cRequests.size(), cRequests.data(), cStats.data() );
368 if( nEntriesComm > 0 )
370 MPI_Request cRequests[2];
371 MPI_Status cStats[2];
374 std::vector< int > dataRowCols( 2 * nEntriesComm );
375 vecRow.Allocate( nEntriesComm );
376 vecCol.Allocate( nEntriesComm );
377 vecS.Allocate( nEntriesComm );
381 mpierr = MPI_Irecv( dataRowCols.data(), 2 * nEntriesComm, MPI_INT, rootProc, iRead * 1000, commW,
385 mpierr = MPI_Irecv( vecS, nEntriesComm, MPI_DOUBLE, rootProc, iRead * 1000 + 1, commW, &cRequests[1] );
389 mpierr = MPI_Waitall( 2, cRequests, cStats );
393 int rindex = 0, cindex = 0;
394 for(
int i = 0; i < nEntriesComm; ++i )
396 const int& vecRowValue = dataRowCols[i * 2];
397 const int& vecColValue = dataRowCols[i * 2 + 1];
399 std::map< int, int >::iterator riter = rowMap.find( vecRowValue );
400 if( riter == rowMap.end() )
402 rowMap[vecRowValue] = rindexMax;
407 rindex = riter->second;
410 std::map< int, int >::iterator citer = colMap.find( vecColValue );
411 if( citer == colMap.end() )
413 colMap[vecColValue] = cindexMax;
418 cindex = citer->second;
421 sparseMatrix( rindex, cindex ) = vecS[i];
429 MPI_Barrier( commW );
434 if(
rank == rootProc )
437 assert( nEntriesRemaining == 0 );
448 printf(
"Rank %d: sparsematrix size = %d x %d\n",
rank, sparseMatrix.GetRows(), sparseMatrix.GetColumns() );
452 int isConsistentP = mapRemap.IsConsistent( dTolerance );
453 if( 0 != isConsistentP )
455 printf(
"Rank %d failed consistency checks...\n",
rank );
466 int isMonotoneP = mapRemap.IsMonotone( dTolerance );
469 mpierr = MPI_Allreduce( &isMonotoneP, &isMonotoneG, 1, MPI_INT, MPI_SUM, commW );
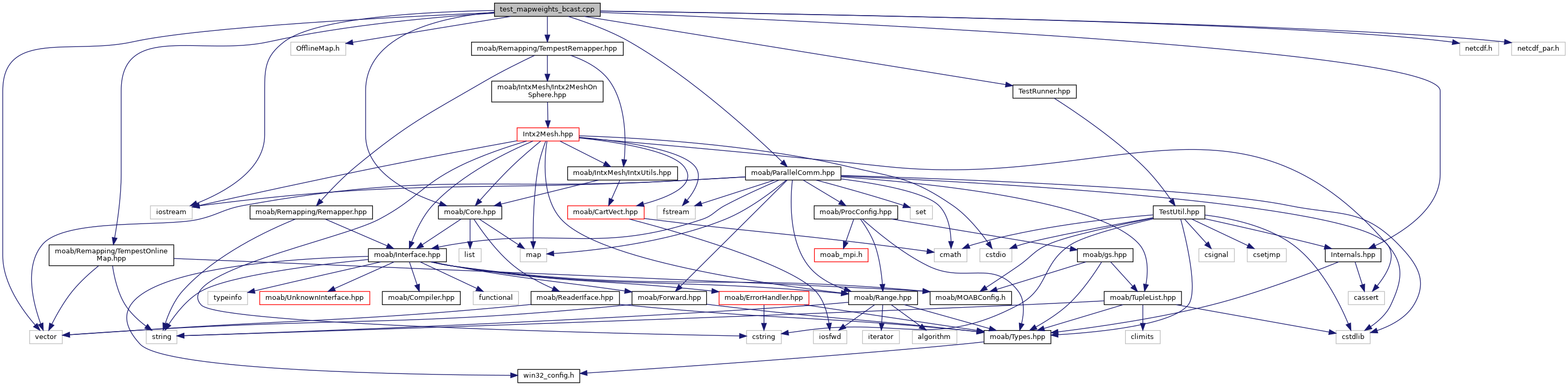
 Include dependency graph for test_mapweights_bcast.cpp:
Include dependency graph for test_mapweights_bcast.cpp: